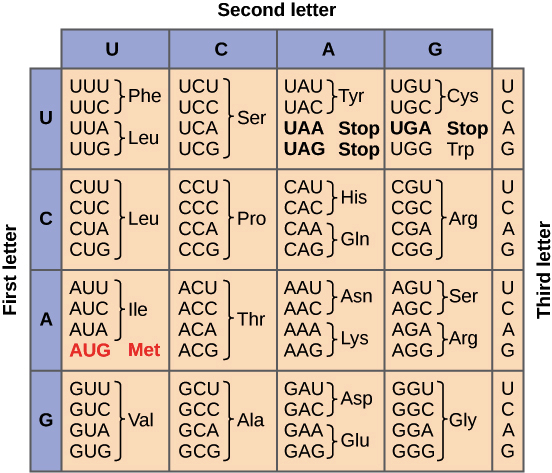
Steps are completed in the nucleus ( Nishikura and De Robertis 1981 Lund and Dahlberg 1998). In vertebrates and higher eukaryotes, the end trimming and splicing TRNA splicing is essential in all eukaryotes examined. Although introns only occur in a subset of tRNAs in different organisms, they are found ubiquitously in eukaryotes, and Intron, all of which must be removed before the tRNA can participate in translation ( Hopper 2013). In the yeast Saccharomyces cerevisiae and other eukaryotes, tRNAs are transcribed by Pol III as pre-tRNAs with a 5′ leader, a 3′ trailer, and in some cases an TRNAs undergo a highly complex maturation process. 2013, 2014) and they must be stable enough to have long half-lives, enabling participation in multiple rounds of translation ( Kadaba et al. 1974 Marck and Grosjean 2002 Valle et al. 2017) they must adopt a conserved structure to uniformly participate in translation, with sufficient flexibility for accommodationĪnd passage through the ribosome ( Kim et al. 2017), and in their anticodon–codon interactions during decoding ( Murphy et al. TRNAs must be selective in charging by their cognate synthetase ( Giege et al. In all organisms, the central role of tRNAs in decoding mRNA during translation places restrictions on the properties of tRNA. Previous Section Next Section INTRODUCTION These results demonstrate that the integrity of the anticodon stem–loop and the efficiency of tRNA splicing are monitored General as it extends to variants of tRNA Tyr(GUA) and tRNA Ser(CGA). Of the intron or by restoration of exon–intron structure, each also resulting in increased tRNA levels. Pathway can contribute substantially to decay of anticodon stem variants, since pre-tRNA decay is largely suppressed by removal 
Mutations that restore exon–intron structure commensurately reduce pre-tRNA accumulation and MPD. Multiple SUP4 οc anticodon stem variants that are subject to MPD each perturb the bulge-helix-bulge structure formed by the anticodon stem–loopĪnd intron, which is important for splicing, resulting in substantial accumulation of end-matured unspliced pre-tRNA as wellĪs pre-tRNA decay. In the anticodon stem are targeted for Met22-dependent pre-tRNA decay (MPD). We provide evidence for a new tRNA quality control pathway in which intron-containing pre-tRNAs with destabilizing mutations RTD is inhibited by mutation of MET22, likely due to accumulation of the Met22 substrate adenosine 3′,5′ bis-phosphate, which inhibits 5′-3′ exonucleases. Or the presence of destabilizing mutations in the acceptor stem, T-stem, or tRNA fold. RTD is catalyzed by theĥ′–3′ exonucleases Xrn1 and Rat1, which act on tRNAs with an exposed 5′ end due to the lack of certain body modifications 
During tRNA maturation in yeast, aberrant pre-tRNAs are targeted for 3′–5′ degradation by the nuclear surveillance pathway,Īnd aberrant mature tRNAs are targeted for 5′–3′ degradation by the rapid tRNA decay (RTD) pathway.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
